Although advances in genetics and genomics reveal numerous disease-associated gene mutations associated with disease, physicians and researchers still wrestle with the tricky challenge of linking those mutations to actual disease-causing processes.
Add this to the growing use of single-cell biology, which detects gene and/or protein expression in each cell at a given moment, and the task becomes even more daunting.
Now, researchers at Cincinnati Children’s Hospital Medical Center report in the journal Nature developing a molecular workflow that leverages single-cell methods to understand the molecular pathways associated with specific patient gene mutations. Scientists first identified a gene mutation that causes the blood disease severe congenital neutropenia (SCN) in children. This mutation was found to block blood cell development leading to the formation of neutrophils.
The study creates a new platform for researchers and clinicians to study the single-cell genomics of a variety of different diseases, which potentially could make genetic-based diagnoses in the clinic more precise and effective, according to the study’s principal investigator, H. Leighton Grimes, Ph.D., in the Division of Immunobiology at Cincinnati Children’s.
“Other than some already well-known causative mutations, determining the difference between important and unimportant DNA changes can be tedious and difficult,” Grimes said. “We can already sequence a child’s genome and link a DNA sequence difference to the disease. However, determining which of these DNA changes is a real disease-causing mutation is difficult, but critical to understanding the molecular mechanisms of a disease, and driving toward a cure.”
Tracking Genetic Mischief
The researchers focused on mutations involving the gene Growth Factor Independent-1 (GFI1). Children with SCN only have immature blood cells in their immune systems, including immature neutrophils. This puts them at high risk for infection and disease.
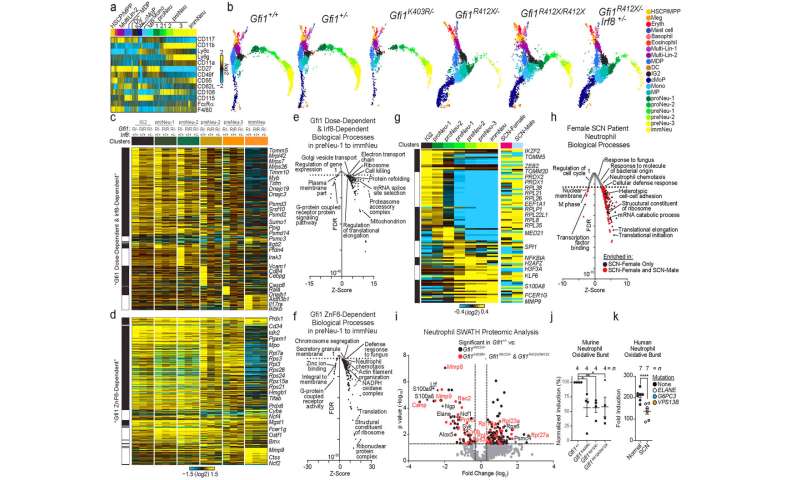
Researchers created genetic models of SCN with mice and human neutrophil cells, which confirmed the disease-causing effect of GFI1 mutations and allowed scientists to determine how the mutant GFI1 affects neutrophil blood cells at each stage of the cell development cycle. That new data allowed them to successfully treat and partially rescue immune cells called neutrophils in their SCN models.
Although genetic manipulations rescued the impact of the GFI1 mutations on neutrophil specification (an initial step of differentiation), the resulting cells were still defective in commitment (a later stage of differentiation where effector functions are programmed). That meant the rescued cells were still unable to function fully as part of the innate immune system, the body’s first line of defense against infection.
The finding underscores the importance of treating each cell type impacted by a disease causing mutation. The new molecular workflow system is expected to help facilitate strategic targeting for therapeutic intervention.
Blending Biology and Supercomputers
The work began within the Division of Human Genetics, using a clinical assay that re-sequenced the genomes of cells from 225 children with SCN. The researchers found DNA sequence alterations in the GFI1 gene. Although some mutations in GFI1 are known to cause neutropenia, many of the GFI1 DNA sequence changes were of unknown clinical significance.
Next, the research team combined computational bioinformatics (lead by co-corresponding author Nathan Salomonis, Ph.D.) with biological experimentation in the laboratory.
To determine whether any of the newly discovered DNA sequence changes in GFI1 could cause neutropenia, researchers created two genetic models. Namely, they genetically introduced the SCN-patient’s GFI1 mutations into the mouse genome, and human induced pluripotent stem cells.
The stem cells can be induced to generate human neutrophil cells.
As the neutrophil cells developed, investigators captured their expressed genes, proteomes and other molecular components in normal and mutant human cells and mice through each stage of the cells’ development cycle. They next employed a new informatics workflow called cellHarmony. This allowed them to detail and compare the downstream target genes and molecular activities of normal and mutant cells at each stage of neutrophil development.
In human cells and mice with mutant GFI1, the mutation triggered a cascade of molecular dysfunction that blocked the development of immature neutrophils. This confirmed the disease causing effect of the GFI1 gene DNA sequence alterations.
That dysfunction was propelled in part by defects in chromatin in the developing mutant cells’ nuclei—the part of the cell that contains its genetic code and DNA. The chromatin in mutant cells remained open and subject to DNA alterations throughout subsequent development stages. This helped scientists identify the cell states most affected by the mutant gene -notably those encompassing neutrophil specification.
The new data allowed researchers to genetically repair neutrophil specification, which in turn controls the number of neutrophils. But repairing neutrophil specification did not automatically repair neutrophil commitment, the development stage where the neutrophils’ innate immune function is programmed.
Children with SCN are regularly treated with a cytokine called Granulocyte colony stimulating factor, a therapy that rescues the numbers of neutrophils produced. But those same children often need to be treated with antibiotics and antifungals to battle various infections.
Grimes said this raises the question of whether cytokine treatment completely repairs the defect, especially since children receiving cytokine therapy for SCN require additional treatment to protect them from recurrent infections. When the researchers analyzed the innate immune function of neutrophils from cytokine-treated SNC patients, they discovered that similar to the mouse cells in their study the cells did not undergo normal commitment and were defective in innate immune function.
Source: Read Full Article
