A team led by researchers at NYU Langone Health’s Perlmutter Cancer Center has identified a gene that drives development of the second most common type of lung carcinoma, offering greater insight into how the disease might be treated.
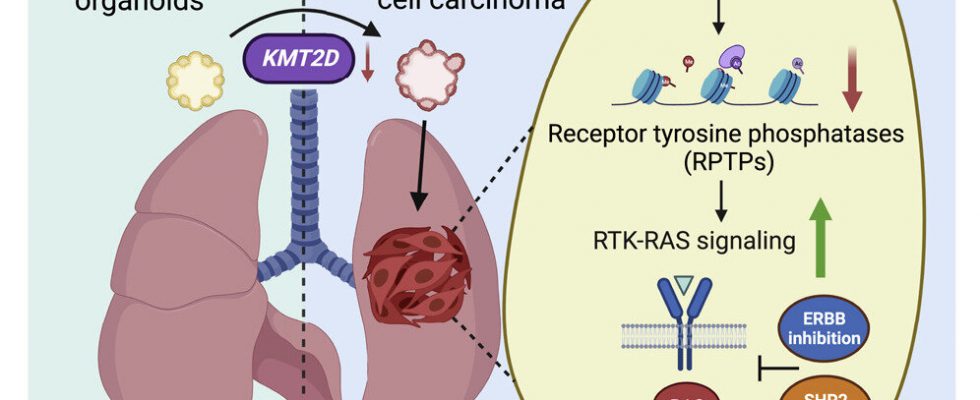
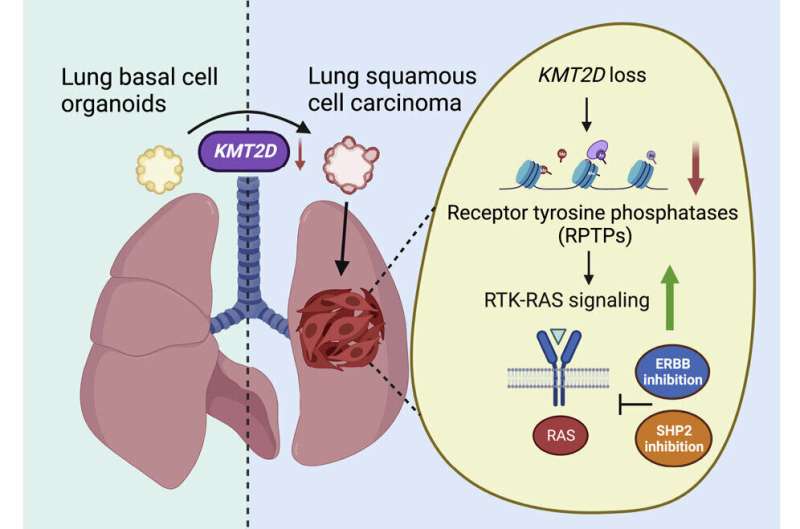
There is currently no approved, targeted, first-line therapy for lung squamous carcinoma (LUSC), a cancer that forms in cell layers lining the organ and is responsible for 20 to 30 percent of lung carcinoma deaths. A new study, published in Cancer Cell, found that deleting a gene called KMT2D caused normal (basal) lung cells grown in complex cultures called organoids to transform into LUSC cells.
According to the study authors, KMT2D regulates the activity of genes that enable the building of protein tyrosine phosphatases, enzymes that restrain the cell-growth-encouraging signals sent through another enzyme set called receptor tyrosine kinases (RTKs). Two RTKs, called EGFR and ERBB2, are known to take part in the abnormal activation of the RTK-RAS signaling pathway, wherein a molecular switch gets “stuck in the on mode,” causing cells to continually multiply as part of cancer.
“Our study identifies KMT2D as a pivotal contributor to the development of lung squamous cancers, and offers vital clues about how to target KMT2D-deficient LUSC,” says co-corresponding author Kwok Kin-Wong, MD, Ph.D., director of the Division of Hematology and Medical Oncology at NYU Langone Health. “The same genetic changes that cause the gene to contribute to cancer also create tumors that are very sensitive to existing drugs that target a related pathway.”
New approaches suggested
The new study confirms prior evidence that the KMT2D gene encodes a protein (a histone methyltransferase) that determines the degree to which the tyrosine phosphatase genes can be accessed by the cellular machinery trying to read them.
Given the better understanding of LUSC mechanisms resulting from the new study, the research team chose to test in study mice a combination of two drugs—SHP2 inhibitor SHP099 and pan-ERBB inhibitor afatinib.
ERBB is made more active by KMT2D signaling flaws, and the enzyme SHP turns up the RTK-KAS pathway, much like EGFR and ERBB2, which are rendered more active by the lack of KMT2D. The team reasoned that experimental drugs designed to inhibit SHP might also counter the effect of KMT2D deficiency when used alongside the ERBB inhibitor.
Indeed, they found that the combination slowed lung tumor growth in mice with LUSC that had been engineered to lack KMT2D, as well as in tumors in mice derived from the human LUSC tumors with KMT2D mutations.
“Multiple SHP2 inhibitors are currently testing in clinical trials, and afatinib is already available,” says co-corresponding author Hua Zhang, MD, Ph.D., formerly an instructor in the Department of Medicine at NYU Langone Health, and now an assistant professor in the Department of Medicine, Division of Hematology and Oncology, at University of Pittsburgh School of Medicine and UPMC Hillman Cancer Center. “Our findings warrant the design of clinical trials that test these therapies in KMT2D-deficient patients with LUSC.”
More information:
Yuanwang Pan et al, KMT2D deficiency drives lung squamous cell carcinoma and hypersensitivity to RTK-RAS inhibition, Cancer Cell (2022). DOI: 10.1016/j.ccell.2022.11.015
Journal information:
Cancer Cell
Source: Read Full Article